On 29 July 2014, HP released the results of a study claiming that 70% of the most commonly used Internet of Things (IoT) devices contained vulnerabilities. Furthermore the devices averaged 25 vulnerabilities per product.
(see http://www8.hp.com/us/en/hp-news/press-release.html?id=1744676 )
So, since Gartner is anticipating something like 26 billion units installed by 2020, there is little doubt that users will be suffering from a myriad of IoT information security and privacy problems well into the next decade. Fortunately, it is still possible to do some things that will reduce the extent of this problem.
While having a complete understanding IoT information security problems is beyond the capability of IoT device users, many will appreciate the value of purchasing devices with a trust-mark. For example when someone buys an electric appliance that has the UL® trust-mark on it, the buyer understands it’s less likely that this product will electrocute someone. Similarly, buyers could come to believe that an IoT security trust-mark will mean that the marked device is less likely to be hacked, less likely to be used to hack other devices, and/or less likely to disclose someone’s personal information.
Devices that come with an IoT security trust-mark would need to meet a standard, and, as with the UL® mark, these standards would need to be verified by an independent third party. Many things could be included and tested according to such standards. Here’s a list of some of the things that might be included along with a brief description of each:
Active Anti-Tamper: FIPS 140 is a NIST standard that describes security for many commercially available cryptographic devices with varying levels of security. The highest level includes physical active anti-tamper capabilities that will cause keys and other critical security parameters to be erased whenever the physical boundaries of the device are penetrated. There are many technologies that can meet these requirements, and many are not that difficult to implement. Similar anti-tamper standards can be applied to IoT devices.
Trusted Boot: Most PCs contain a device called a Trusted Platform Module (TPM). This device can be used to help ensure that the code executed while booting has not changed from one boot to the next. If the boot code has changed from an authorized version, the TPM makes it possible for other devices to stop trusting the changed system. Trusted IoT devices should have hardware for verifying the device has booted into a trusted state.
Removeable Power: Many cell phones today, have removable batteries, and many people have realized that this is a strong security feature. By removing the batteries from a cell phone, a user can be relatively certain that he has disabled any spyware that might be running on the phone, spyware that might be listening to the user’s conversations or reporting on the user’s location. A user of a trusted IoT device should have the ability to stop trusting that device by removing the power source.
Independent User Control of Physical I/O Channels: Similarly, a user, not wanting to completely disable his device, might wish to be sure certain I/O functions are not activated. For example, the user may want to disable the camera function, the GPS function and the microphone function while retaining the ability to listen to music. By providing hardwired switches certified to disable specific hardware I/O function, a user can rest assured that these functions won’t be secretly activated by some malware lurking inside the trusted IoT device.
Host Based Intrusion Detection: For several years now, host based intrusion detection software has been available for desktop machines and servers. It is time to recognize that IoT devices are hosts too. There should be software running on the trusted IoT device so that one can detect when that trust is no longer appropriate.
Automatic Security Patching: Today, the time between the release of a critical security patch and the release of malware that exploits the associated vulnerability can be measured in hours. The reality of the present situation is that the existence of a critical security patch means your system is already broken. Consequently, the automated application of security patches is necessary for desktops and servers. Automated security patching for trusted IoT devices will also be necessary.
Independent Software Security Verification: To a certain extent, trusting a software companies to develop secure code is like trusting a fox to guard a hen house. This is because the pressures on software developers to make marketing windows, to release code and to get paid frequently overpower discussions about the appropriate levels of security needed for operating the end products safely. The resulting security problems are then left for others to solve. Because of this, various information security standards depend on independent software security verification. While this can be expensive, free services like “The SWAMP”
( https://continuousassurance.org/about-us/ ) offer the hope that independent software security verification can be done cheaply enough to motivate standardization.
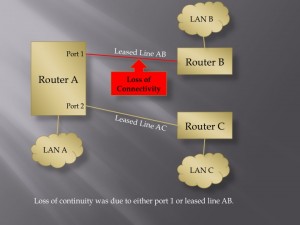
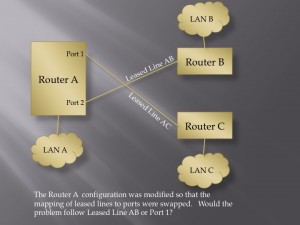
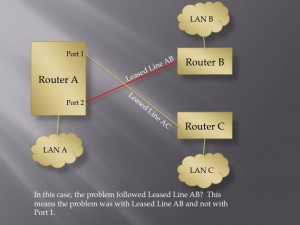
User Defined Trust Relationships: When an IoT device enters a home, there may be very good reasons why it will need to communicate with other devices inside or outside of that home. That does not mean that the new device should have the ability to communicate with all other devices. Consider the recent Target hack. The point of sale terminals were attacked by first gaining access to a system used to manage heating, ventilation and air conditioning. Likewise, it might not make sense for your home’s air conditioning system to be able to talk with your home’s electric door locks. It seems that giving users an easy way to manage what systems are allowed to talk with other systems could help quite a bit here. How to do this effectively may take some creativity, but one could imagine users having a tool, perhaps a wand that they could tap on one device and then tap on another device, to establish or dissolve the trust relationships between devices.
Recently, on 10 September 2014, The International Workshop on Secure Internet of Things (SIOT 2014, see http://siot-workshop.org/ ) conducted its meeting in Wroclaw, Poland. This was only the third such workshop. So, SIOT standardization is still far from being where it needs to be. What will actually go into a set of IoT Security standards is not yet known. Likewise, an IoT Security Trustmark is not yet available. Hopefully, some of the ideas suggested above will start to find their way into trusted IoT devices. If not, we can surely expect the same sorts of security problems that have plague our PCs and web servers, to appear all over again in the Internet of Things.